-
New paper! Our latest study reveals the individual-to-individual variation of PGS accuracy across the genetic-ancestry continuum, even in traditionally defined "homogenous" populations! (Ding et al 2023). Read the Research Briefing, UCLA Health News and twitter thread for more details.May 2023
-
New paper! Our recent analysis of individuals of admixed genetic ancestries suggests that complex trait causal variant effect sizes are largely similar across ancestries, and discusses the implication for the study of these and other diverse populations. (Hou et al 2023). Read the Research Briefing by Elizabeth G. Atkinson and twitter thread for more details.March 2023
-
New paper! We proposed a general framework to quantify the uncertainty of an individual's PRS estimate using Bayesian approach. Analysis of 13 complex traits from UK Biobank show large credible interval in individual PRS estimates which impacts the interpretation of PRS-based risk stratification (Ding*, Hou* et al. 2021). Read the twitter thread for more details.Dec 2021
-
New paper! We argue that Tractor reduces GWAS power in admixed pops in realistic settings and we recommend caution when using it for GWAS in admixed populations (Hou 2021). Read the twitter thread for more details.Dec 2021
-
New paper! We propose a statistical framework to estimate regional polygenicity for a complex trait using marginal effect sizes from GWAS and LD information. We find that SNP-heritability is proportional to polygenicity both on the genome-wide and regional scale, suggesting that the observed differences in heritability across traits stem from differences in the underlying number of causal SNPs(Johnson 2021).Oct 2021
-
Congratulations to Eva for graduating with a BS in Computational and Systems Biology with highest honors!June 15 2021
-
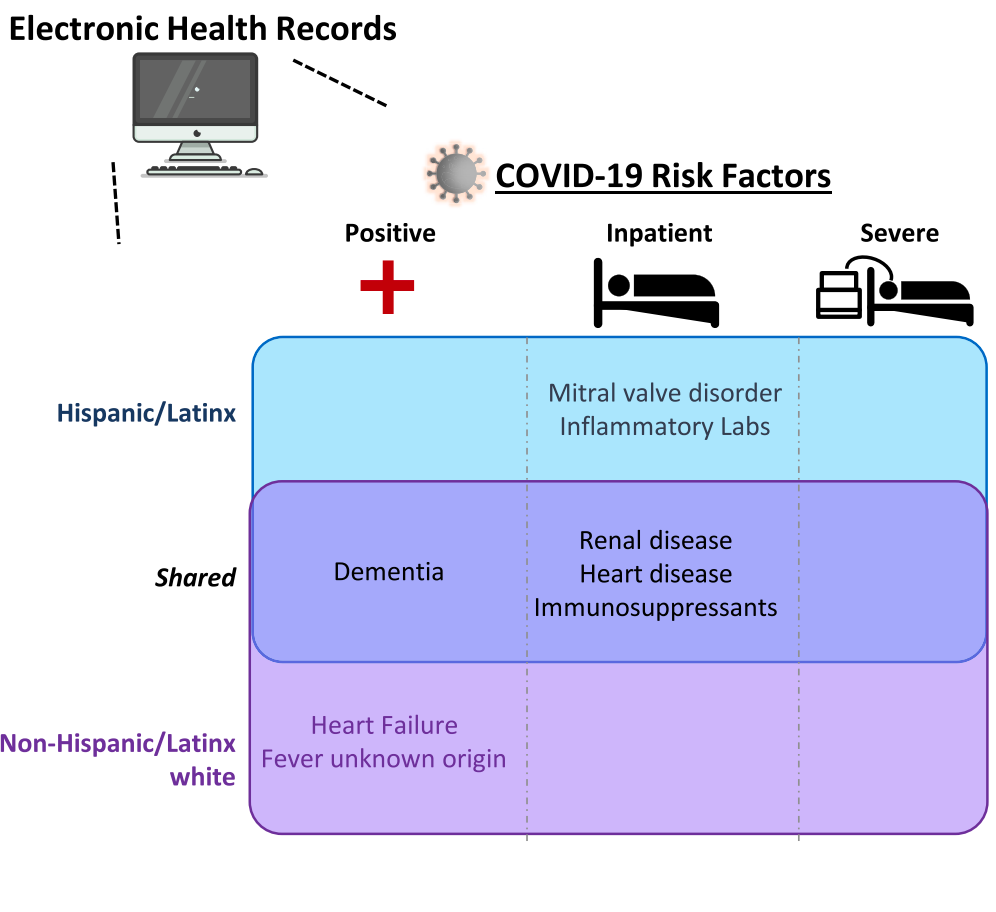
New paper! We find a higher risk of COVID-19 among Hispanics/Latinx patients from UCLA health system and identify risk factors that are specific to this underrepresented population (Chang*, Ding*, Freund*, Johnson*, Schwarz* et al. 2021)March 2021
-
Read Bogdan's thoughts on diversity in genetic studies and its role in demystifying COVID-19.October 2020
-
Congratulations to Dr. Freund for graduating with her PhD in Human Genetics! Malika is moving on to pursue a Master’s in Genetic Counseling at Stanford University. She will be missed but we can’t wait to see what she accomplishes next!
 June 30 2020
June 30 2020 -
New Paper! Our novel method PESCA estimates the numbers of causal genetic variants shared across ancestral populations (Shi*, Burch* et al. 2020). Check out our blog post. which summarizes the questions that motivated us and our main findings.May 2020
-
Congratulations to Dr. Malika Kumar Freund! The first online defense in Bogdan Lab’s history attracted an audience of more than 80 people!
 March 20 2020
March 20 2020 -
-
Tommer gets grilled during the poster session.
 Oct 2019
Oct 2019 -
-
We'll miss you, Annique! Thanks for a fun summer in the lab!
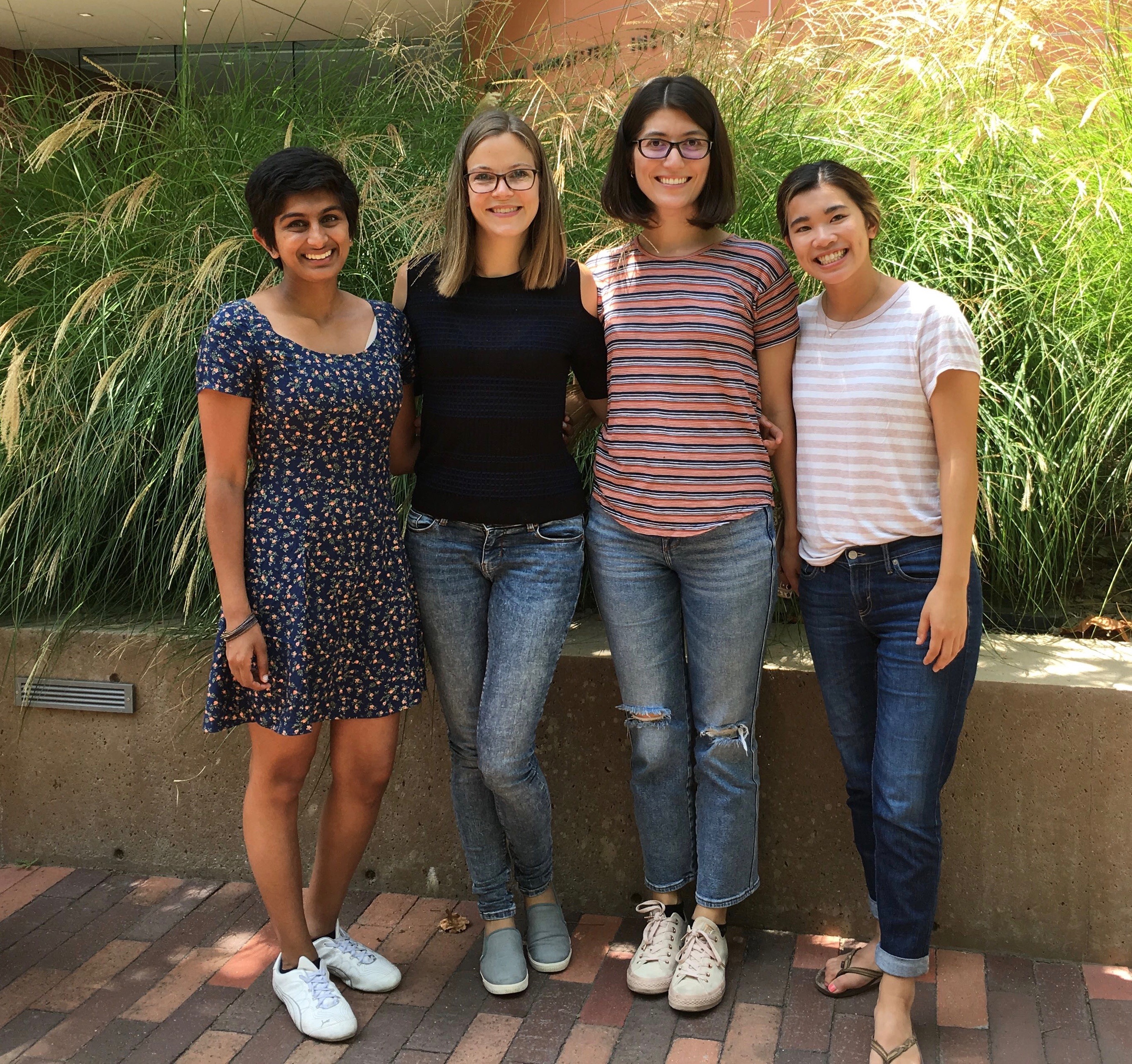 Aug 2019
Aug 2019 -
Gary and Hugo present their posters at the BIG Summer Symposium. Congrats to Gary for taking home a Best Poster Award!
 Aug 2019
Aug 2019 -
New Paper! Our latest work features robust SNP-heritablity estimation in biobank-scale data (Hou*, Burch* et al. 2019).Jul 2019
-
Congratulations to Kangcheng, who graduated from Zhejiang University with a B.Eng in Computer Science! He will be joining the Bioinformatics PhD program at UCLA this fall.July 2019
-
New Paper! Our recent collaborative work examines the enrichment of breast and prostate cancer GWAS associations in regulatory regions across tissue types (Chen et al. 2019).Jun 2019
-
New Paper! In collaboration with the Ophoff Lab, we identify a novel risk locus for Dupuytren's disease and explore shared genetic etiology with BMI (Major et al. 2019).May 2019
-
Read our feature story in the newsrooms of the UCLA Jonsson Comprehensive Cancer Center and Cedars-Sinai!May 2019
-
Our latest work, in collaboration with Cedars-Sinai Cancer and Dana-Farber Cancer Institute, identifies 34 genes associated with an increased risk for developing the earliest stages of ovarian cancer.(Gusev et al. 2019).May 2019
-
Ruthie and Bogdan are giving talks at the 2019 Biology of Genomes in May!
Ruthie: "Dissecting the genetic architecture of complex traits through local polygenicity using summary statistics from genome-wide association studies"
Bogdan: "Accurate estimation of SNP-heritability from biobank-scale date irrespective of genetic architecture"
April 2019 -
Two publications in the latest issue of Nature Genetics! FOCUS: fine-mapping TWAS to find causal genes (Mancuso et al. 2019).March 2019
-
Also in the latest issue of Nature Genetics, in collaboration with many, we present a perspective on the usage and interpretation of TWAS as a tool to find causal genes (Wainberg et al. 2019).March 2019
-
Congratulations to Nick, who will be joining the Center for Genetic Epidemiology and Department of Preventive Medicine at USC as a tenure-track assistant professor next month!March 2019
-
New Paper! An examination of 296,215 cases across 6 solid cancer types highlights some shared common germline genetic basis, suggesting that some cancers could be studied together to better understand the biological pathways that lead to cancer (Jiang et al. 2019).January 2019
-
New Paper! Our latest work with the Price Lab uses polygenic functional enrichment to incorporate coding, conserved, regulatory, and LD-related genomic annotations into association analyses (Kichaev et al. 2019).January 2019
-
Blog update: Read Ruthie's 7 Tips for making presentations, a compilation of quick tips that our lab finds useful when putting together presentations!December 2018
-
Arun presents his work on tissue-wise genetic subtyping of complex traits in the Genetics of Cardiometabolic Traits session at ASHG!
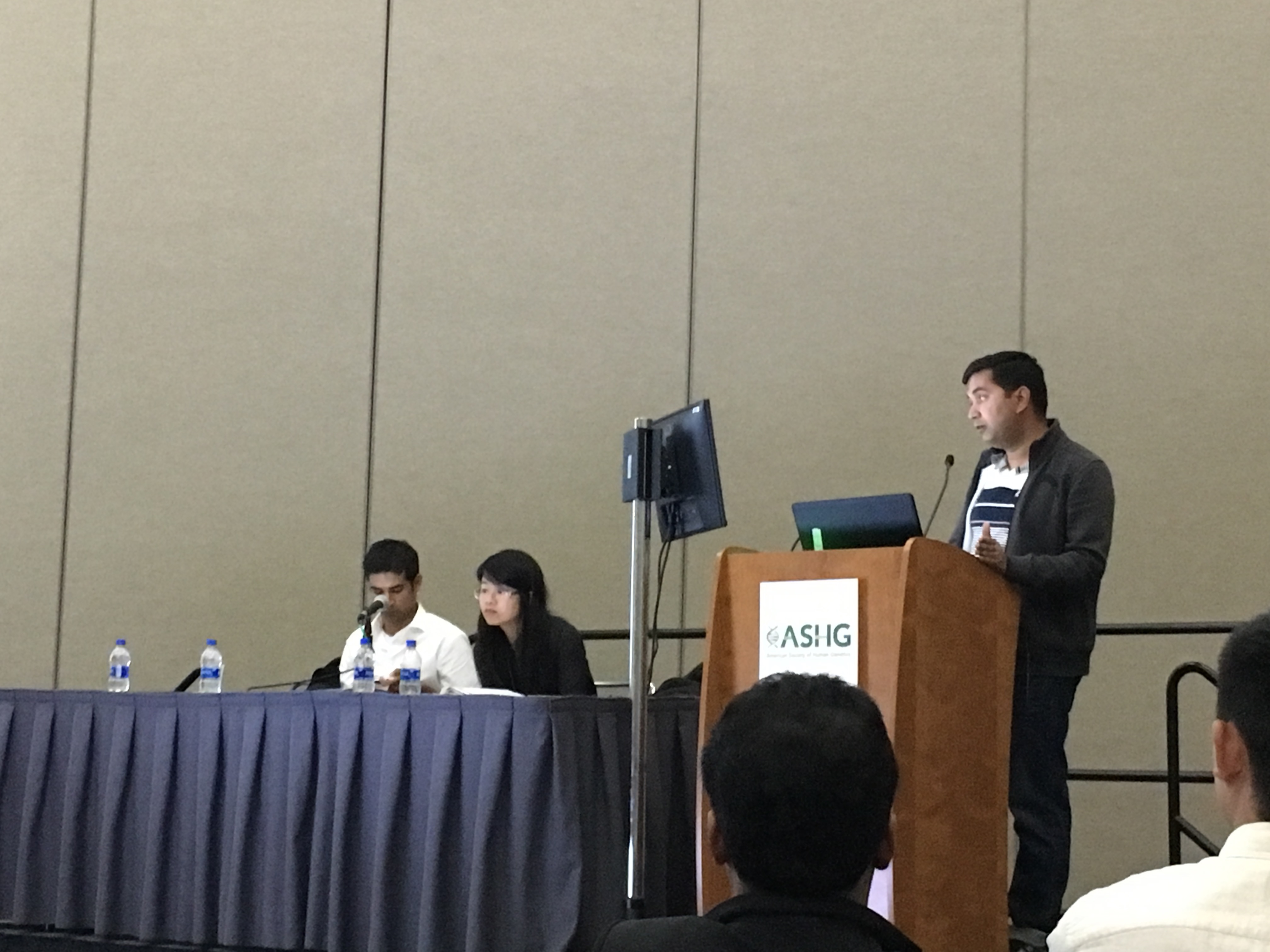 October 2018
October 2018 -
The lab attends ASHG 2018 in San Diego!
 October 2018
October 2018 -
New Paper with the Arboleda Lab! Our quantification of the shared genetic basis between complex traits and Mendelian disorders is now published in AJHG (Freund et al. 2018).October 2018
-
New Paper! Our latest TWAS for prostate cancer risk is now published in Nature Communications (Mancuso et al. 2018).October 2018
-
Kangcheng presents his poster on fixed effects models for heritability estimation at the CSST Summer Poster Session!
 Sept 2018
Sept 2018 -
Arun's abstract on tissue-wise subtyping of complex traits is selected for a platform presentation at the American Society of Human Genetics Annual Meeting!August 2018
-
Congratulations to Dr. Megan Roytman for successfully defending her thesis!
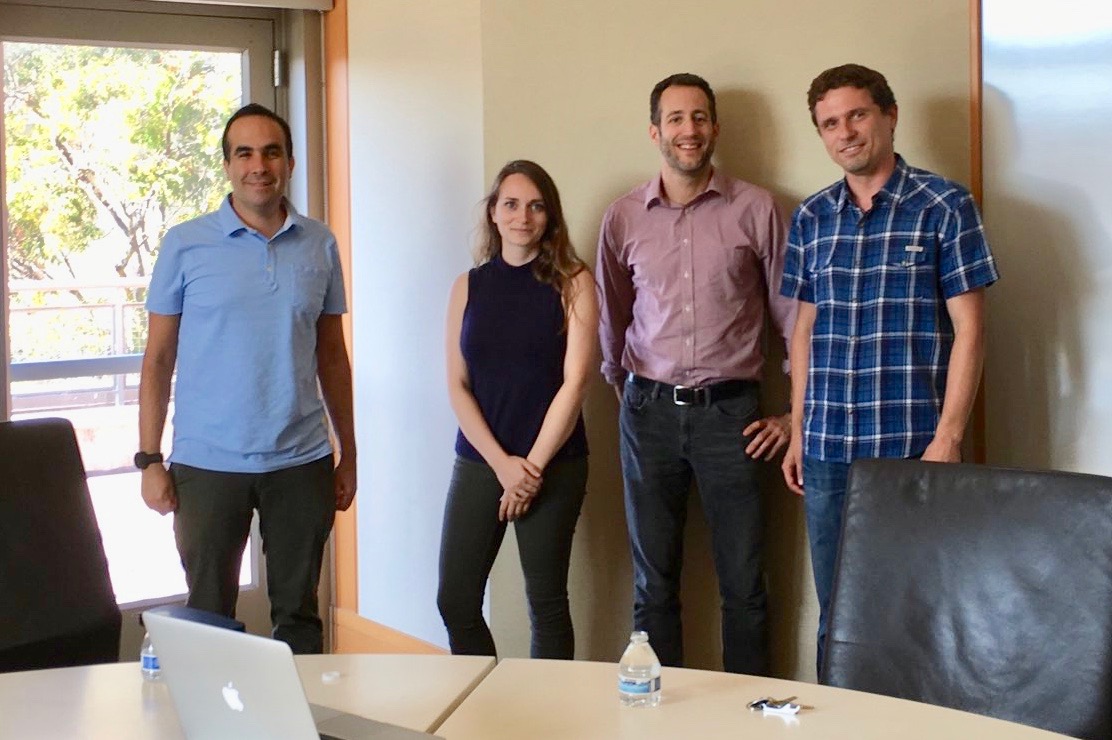 July 2018
July 2018 -
Ruthie gives a talk on the UNITY framework at the Intelligent Systems for Molecular Biology annual conference in Chicago!
 July 2018
July 2018 -
-
It's graduation day -- Drs. Kichaev and Shi await their diplomas!
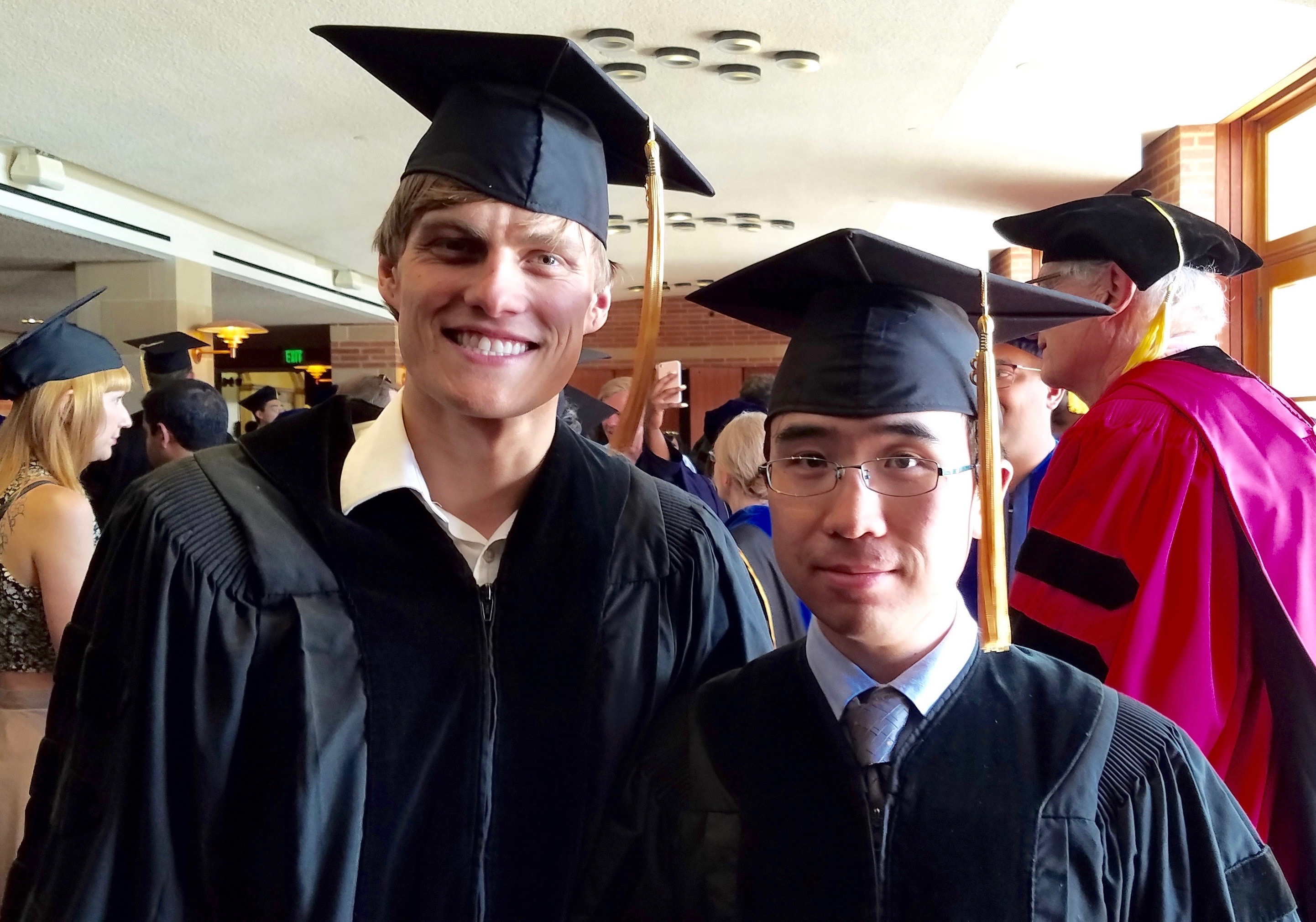 June 2018
June 2018 -
TWAS hub is now live, offering searchable access to TWAS results from hundreds of complex traits and dozens of expression studies!June 2018
-
Nick and Malika earn fellowships at UCLA in the Neurobehavioral Genetics Training Grant and Biomedical Big Data Training Grant, respectively!June 2018
-
Congratulations on a successful PhD defense to our newly-minted Dr. Huwenbo Shi!
 May 2018
May 2018 -
Malika's abstract is selected for an oral presentation at the UCLA Annual Joint Research Symposium!April 2018
-
Kathy earns a fellowship in the Genomic Analysis and Interpretation Training Program at UCLA!April 2018
-
When the fire alarm goes off during your PhD defense, you have to improvise! Congratulations to our own Dr. Gleb Kichaev!
 April 2018
April 2018 -
-
-
Blog update: Check out our latest post on Explaining missing heritability using Gaussian process regression by Huwenbo!February 2018
-
Blog update: Working with GWAS summary data? Read Huwenbo's tips for formatting and quality control to avoid pitfalls in post-GWAS analyses!February 2018
-
Blog update: Read Ruthie's post about CANVIS, a command line fine-mapping tool for visualizing fine-mapping studies.January 2018
-
-
-
The lab attends ASHG 2017 in Orlando!
 October 2017
October 2017 -
Congratulations to Malika for winning the Stephen D. Cederbaum ASHG Abstract Travel Award 2017!October 2017
-
Congratulations to Ruthie and Page for winning the UCLA Dean’s Prize for Excellence at the undergraduate Research Poster Day 2017!
 June 2017
June 2017 -
Gleb wins Cotterman award for best trainee paper in AJHG 2015!
 October 2016
October 2016